 Alfalfa Gene Editing Database
Alfalfa Gene Editing Database
 Alfalfa Gene Editing Database
Alfalfa Gene Editing Database
Alfalfa ( Medicago sativaL.), with high nutritional value and good palatability,
is a perennial herb of the legume family Alfalfa and one of the most widely distributed and longest cultivated forages in the world,
an important forage resource for livestock development and known as the "king of forages".
Genomic data of alfalfa have been published and are important to provide valuable genetic resources for researchers and breeders to improve the traits of this species.
For this purpose, we have developed a comprehensive and easy-to-use web-based database, AlfalfaGEDB.
In this database, genomic and transcriptomic data of two cultivars of alfalfa
(XinJiangDaYe and ZhongmuNo.1)
were collected and analyzed. Based on the development and application of CRISPR gene editing technology,
we made predictions of sgRNA targets.
To better explore the functions of alfalfa,
this database also provides some bioinformatics tools: BLAST, Heatmap, Network, TFs/TRs, Protein Kinases...
AlfalfaGEDB contains seven modules: Home, Genome, BLAST , Search, Tools, Download and Help pages. The framework of the whole system is shown below.
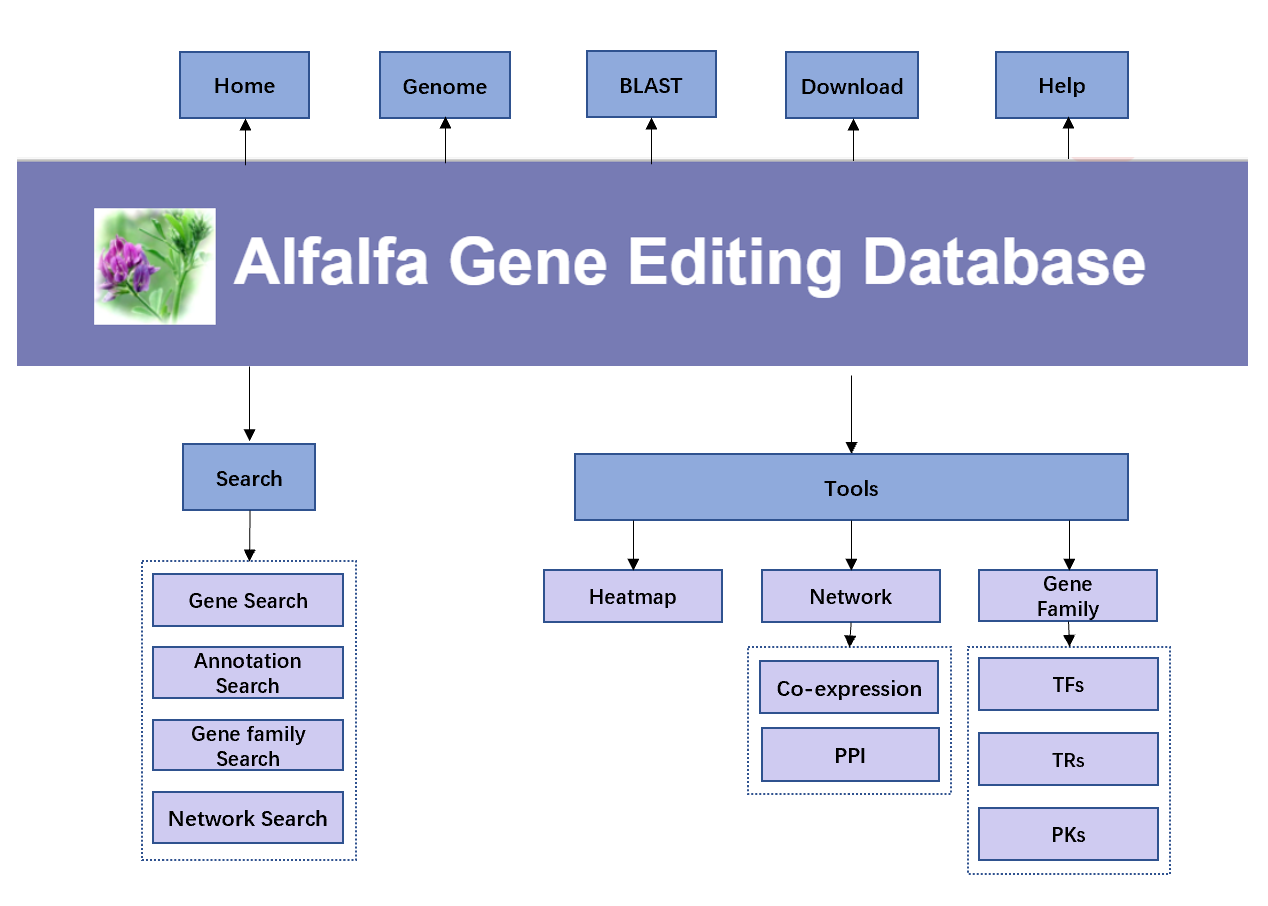
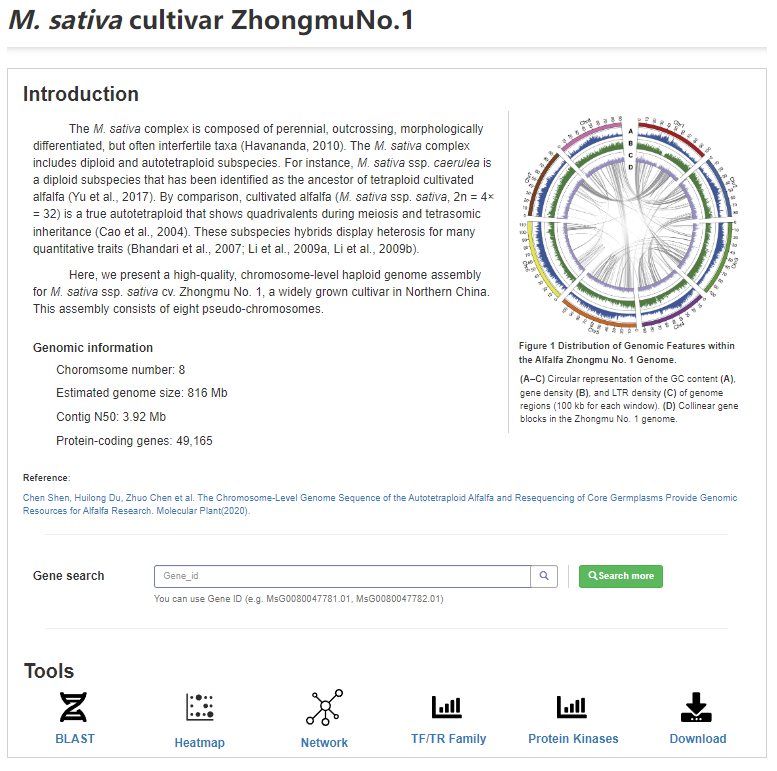
On the Genome page, users can not only get an introduction to the Genome, but also independently browse and query the detailed information of related genes, including location information, sequence information, annotation information and sgRNA prediction information. In addition, quick links to some of the tools are provided.
The BLAST database constructed by AlfalfaGEDB contains mainly the nucleic acid database and protein database of alfalfa. To perform the alignment, the user can select four alignment programs, Blastn, Blastp, Blastx and Tblastn. In addition to the default parameters, user-defined parameters are also allowed: E-value Word size, etc.

The search page includes four sections: search by gene IDs, search by gene annotations, search by gene family classification and search by networks.
1. Search by gene ID. After selecting the reference genome, the user can enter the gene ID to query the details.
2. Querying by annotation information. The user selects the genome and the specific annotation database and enters the annotation to ID or annotation information to get all the genes annotated to that class. Similarly, users can jump to the detailed gene description page by clicking on the gene ID.
3. Query for gene family classification. By entering a gene family and selecting the reference genome, the user can obtain all the genes contained in that gene family
4. Query network data based on gene ID. Users can query the protein interactions network data or co-expression network data by entering the gene ID.

We provide several functions on the tools page for users to choose from:
1. Heatmap: visualize the gene expression data of alfalfa as a heatmap based on the gene sequence given by the user using Clustergrammer (Fernandez et al., 2017).
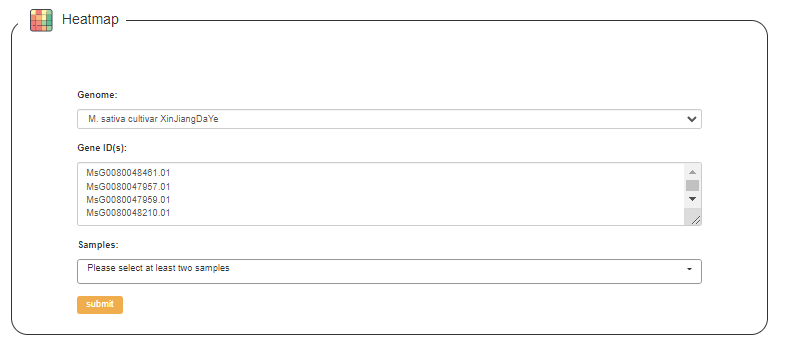
2. Network: generate a biological network by selecting the type of query you want among the co-expression network and protein interactions network based on the gene set provided by the user.

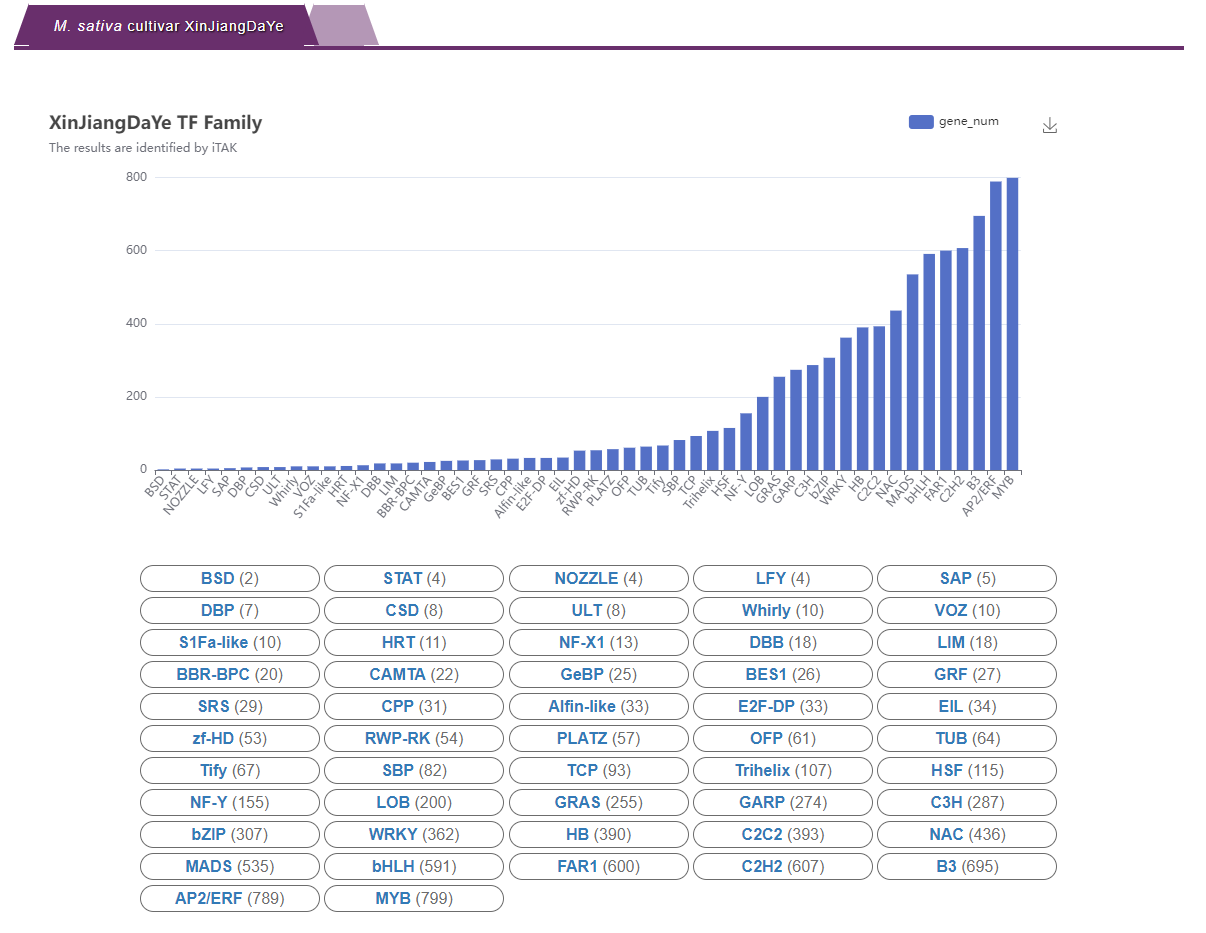
3. Gene family classification: Provide gene family prediction, including TFs, TRs and PKs. The number of genes predicted for each family is shown in a bar graph, and the specific text information is displayed. Users will get the corresponding detailed information after clicking on the bar or text.
Users can quickly obtain the genomic data and annotation information they want to download on the download page.
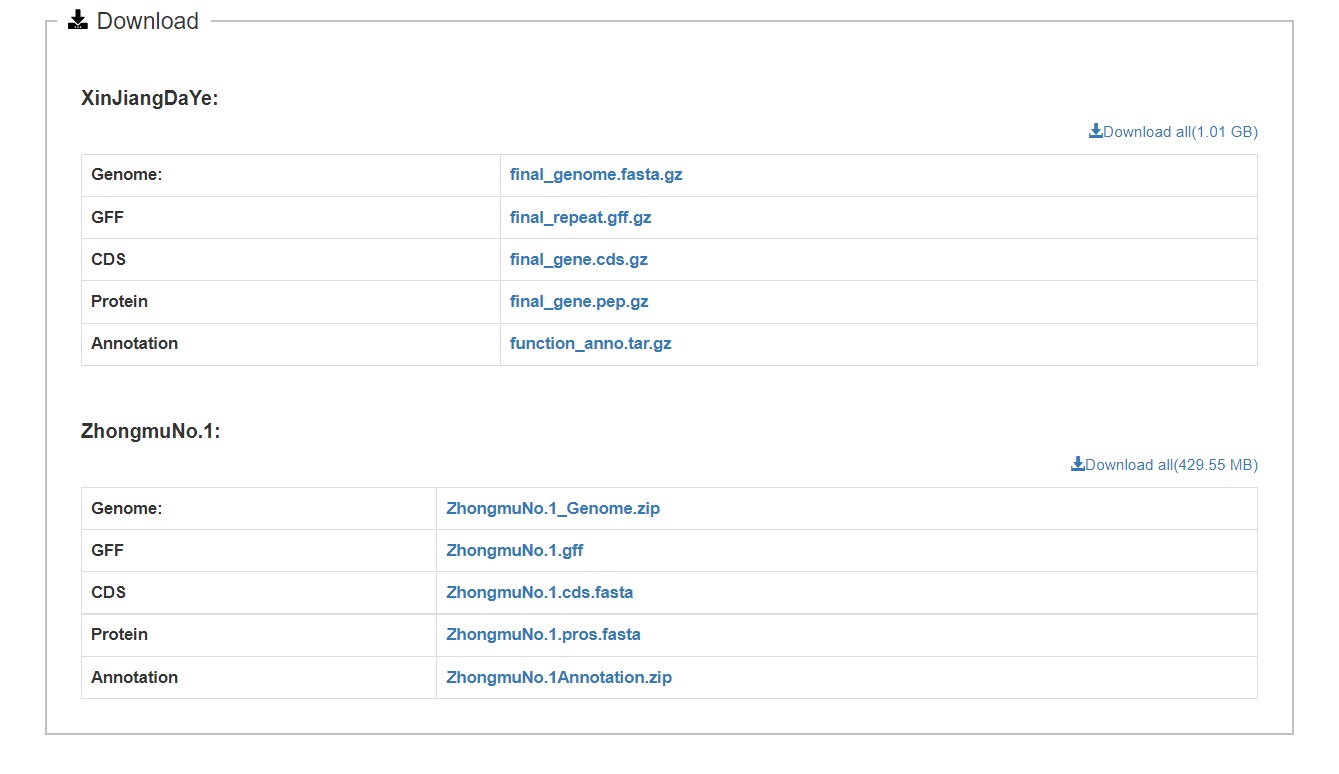
Genomic data
AlfalfaGEDB includes genomic data for two cultivars of alfalfa:
XinJiangDaYe genome data collected from
Allele-aware chromosome-level genome assembly and efficient transgene-free genome editing for the autotetraploid cultivated alfalfa.
ZhongmuNo.1 genome data collected from
The Chromosome-Level Genome Sequence of the Autotetraploid Alfalfa and Resequencing of Core Germplasms Provide Genomic Resources for Alfalfa Research.
transcriptome data
All publicly available transcriptome datasets are retrievable from the NCBI SRA dataset. The deadline for data collection is January 24, 2022.
| Study Accession | Run Accession | Tissue | Condition | Abbreviation |
|---|---|---|---|---|
| PRJNA596791 | SRR10742428 | leaf | Flood-stressed | AAC-TruFL1 |
| PRJNA596791 | SRR10742429 | leaf | Control | AAC-TruCon3 |
| PRJNA596791 | SRR10742430 | leaf | Control | AAC-TruCon2 |
| PRJNA596791 | SRR10742431 | leaf | Control | AAC-TruCon1 |
| PRJNA596791 | SRR10742432 | leaf | Flood-stressed | WT_FL3 |
| PRJNA596791 | SRR10742433 | leaf | Flood-stressed | WT_FL2 |
| PRJNA596791 | SRR10742434 | leaf | Flood-stressed | WT_FL1 |
| PRJNA596791 | SRR10742435 | leaf | Flood-stressed | SPL13_FL3 |
| PRJNA596791 | SRR10742436 | leaf | Flood-stressed | SPL13_FL2 |
| PRJNA596791 | SRR10742437 | leaf | Flood-stressed | SPL13_FL1 |
| PRJNA596791 | SRR10742438 | leaf | Control | SPL13_Con3 |
| PRJNA596791 | SRR10742439 | leaf | Control | SPL13_Con2 |
| PRJNA596791 | SRR10742440 | leaf | Control | SPL13_Con1 |
| PRJNA596791 | SRR10742441 | leaf | Flood-stressed | A8_FL3 |
| PRJNA596791 | SRR10742442 | leaf | Flood-stressed | A8_FL2 |
| PRJNA596791 | SRR10742443 | leaf | Flood-stressed | A8_FL1 |
| PRJNA596791 | SRR10742444 | leaf | Control | A8_Con3 |
| PRJNA596791 | SRR10742445 | leaf | Control | WT_Con3 |
| PRJNA596791 | SRR10742446 | leaf | Control | A8_Con2_R1 |
| PRJNA596791 | SRR10742447 | leaf | Control | A8_Con1 |
| PRJNA596791 | SRR10742448 | leaf | Flood-stressed | AC-CarFL3 |
| PRJNA596791 | SRR10742449 | leaf | Flood-stressed | AC-CarFL2 |
| PRJNA596791 | SRR10742450 | leaf | Flood-stressed | AC-CarFL1 |
| PRJNA596791 | SRR10742451 | leaf | Control | AC-CarCon3 |
| PRJNA596791 | SRR10742452 | leaf | Control | AC-CarCon2 |
| PRJNA596791 | SRR10742453 | leaf | Control | AC-CarCon1 |
| PRJNA596791 | SRR10742454 | leaf | Flood-stressed | AAC-TruFL3 |
| PRJNA596791 | SRR10742455 | leaf | Flood-stressed | AAC-TruFL2 |
| PRJNA596791 | SRR10742456 | leaf | Control | WT_Con2 |
| PRJNA596791 | SRR10742457 | leaf | Control | WT_Con1 |
| PRJNA598830 | SRR10883588 | basal-stem | Drought | EV-Stem-trt1 |
| PRJNA598830 | SRR10883589 | basal-stem | Control | EV-Stem-Con3 |
| PRJNA598830 | SRR10883590 | basal-stem | Control | EV-Stem-Con2 |
| PRJNA598830 | SRR10883591 | basal-stem | Control | EV-Stem-Con1 |
| PRJNA598830 | SRR10883592 | leaf | Drought | EV-Leaf-trt3 |
| PRJNA598830 | SRR10883593 | leaf | Drought | EV-Leaf-trt2 |
| PRJNA598830 | SRR10883594 | root tip | Drought | SPL13-root-trt3 |
| PRJNA598830 | SRR10883595 | root tip | Drought | SPL13-root-trt2 |
| PRJNA598830 | SRR10883596 | root tip | Drought | SPL13-root-trt1 |
| PRJNA598830 | SRR10883597 | root tip | Control | SPL13-root-Con3 |
| PRJNA598830 | SRR10883598 | root tip | Control | SPL13-root-Con2 |
| PRJNA598830 | SRR10883599 | root tip | Control | SPL13-root-Con1 |
| PRJNA598830 | SRR10883600 | leaf | Drought | EV-Leaf-trt1 |
| PRJNA598830 | SRR10883601 | basal-stem | Drought | SPL13-Stem-trt3 |
| PRJNA598830 | SRR10883602 | basal-stem | Drought | SPL13-Stem-trt2 |
| PRJNA598830 | SRR10883603 | basal-stem | Drought | SPL13-Stem-trt1 |
| PRJNA598830 | SRR10883604 | basal-stem | Control | SPL13-Stem-Con3 |
| PRJNA598830 | SRR10883605 | basal-stem | Control | SPL13-Stem-Con2 |
| PRJNA598830 | SRR10883606 | basal-stem | Control | SPL13-Stem-Con1 |
| PRJNA598830 | SRR10883607 | leaf | Drought | SPL13-Leaf-trt3 |
| PRJNA598830 | SRR10883608 | leaf | Drought | SPL13-Leaf-trt2 |
| PRJNA598830 | SRR10883609 | leaf | Drought | SPL13-Leaf-trt1 |
| PRJNA598830 | SRR10883610 | leaf | Control | SPL13-Leaf-Con3 |
| PRJNA598830 | SRR10883611 | leaf | Control | EV-Leaf-Con3 |
| PRJNA598830 | SRR10883612 | leaf | Control | SPL13-Leaf-Con2 |
| PRJNA598830 | SRR10883613 | leaf | Control | SPL13-Leaf-Con1 |
| PRJNA598830 | SRR10883614 | root tip | Drought | EV-root-trt3 |
| PRJNA598830 | SRR10883615 | root tip | Drought | EV-root-trt2 |
| PRJNA598830 | SRR10883616 | root tip | Drought | EV-root-trt1 |
| PRJNA598830 | SRR10883617 | root tip | Control | EV-root-Con3 |
| PRJNA598830 | SRR10883618 | root tip | Control | EV-root-Con2 |
| PRJNA598830 | SRR10883619 | root tip | Control | EV-root-Con1 |
| PRJNA598830 | SRR10883620 | basal-stem | Control | EV-Stem-trt3 |
| PRJNA598830 | SRR10883621 | basal-stem | Drought | EV-Stem-trt2 |
| PRJNA598830 | SRR10883622 | leaf | Control | EV-Leaf-Con2 |
| PRJNA598830 | SRR10883623 | leaf | Control | EV-Leaf-Con1 |
| PRJNA622603 | SRR11467825 | leaf | Caoyuan No.2 with thrips infestation | ST_1 |
| PRJNA622603 | SRR11467826 | leaf | Caoyuan No.2 without thrips infestation | SCK_3 |
| PRJNA622603 | SRR11467827 | leaf | Caoyuan No.2 without thrips infestation | SCK_2 |
| PRJNA622603 | SRR11467828 | leaf | Caoyuan No.2 without thrips infestation | SCK_1 |
| PRJNA622603 | SRR11467829 | leaf | Caoyuan No.4 with thrips infestation | RT_3 |
| PRJNA622603 | SRR11467830 | leaf | Caoyuan No.4 with thrips infestation | RT_2 |
| PRJNA622603 | SRR11467831 | leaf | Caoyuan No.4 with thrips infestation | RT_1 |
| PRJNA622603 | SRR11467832 | leaf | Caoyuan No.4 without thrips infestation | RCK_3 |
| PRJNA622603 | SRR11467833 | leaf | Caoyuan No.2 with thrips infestation | ST_3 |
| PRJNA622603 | SRR11467834 | leaf | Caoyuan No.2 with thrips infestation | ST_2 |
| PRJNA622603 | SRR11467835 | leaf | Caoyuan No.4 without thrips infestation | RCK_2 |
| PRJNA622603 | SRR11467836 | leaf | Caoyuan No.4 without thrips infestation | RCK_1 |
| PRJNA414493 | SRR6227642 | leaf | control | CZM-2 |
| PRJNA414493 | SRR6227643 | leaf | control | CZM-3 |
| PRJNA414493 | SRR6227644 | leaf | 0.5M NaCl | TXJ-2 |
| PRJNA414493 | SRR6227645 | leaf | 0.5M NaCl | TXJ-3 |
| PRJNA414493 | SRR6227646 | leaf | 0.5M NaCl | TZM-1 |
| PRJNA414493 | SRR6227647 | leaf | 0.5M NaCl | TZM-2 |
| PRJNA414493 | SRR6227648 | leaf | control | CXJ-1 |
| PRJNA414493 | SRR6227649 | leaf | control | CXJ-2 |
| PRJNA414493 | SRR6227650 | leaf | control | CXJ-3 |
| PRJNA414493 | SRR6227651 | leaf | control | CZM-1 |
| PRJNA414493 | SRR6227652 | leaf | 0.5M NaCl | TZM-3 |
| PRJNA414493 | SRR6227653 | leaf | 0.5M NaCl | TXJ-1 |
| PRJNA525327 | SRR8667732 | root | drought-resistant | ZT2 |
| PRJNA525327 | SRR8667733 | root | Drought-sensitive | GC3 |
| PRJNA525327 | SRR8667734 | root | Drought-sensitive | GT1 |
| PRJNA525327 | SRR8667735 | root | drought-resistant | ZT3 |
| PRJNA525327 | SRR8667736 | root | drought-resistant | ZC1 |
| PRJNA525327 | SRR8667737 | root | drought-resistant | ZC2 |
| PRJNA525327 | SRR8667738 | root | drought-resistant | ZC3 |
| PRJNA525327 | SRR8667739 | root | drought-resistant | ZT1 |
| PRJNA525327 | SRR8667740 | root | Drought-sensitive | GT2 |
| PRJNA525327 | SRR8667741 | root | Drought-sensitive | GT3 |
| PRJNA525327 | SRR8667742 | root | Drought-sensitive | GC1 |
| PRJNA525327 | SRR8667743 | root | Drought-sensitive | GC2 |
| PRJNA540215 | SRR9026566 | leaf primodia | palm1 mutant leaf primodia sample 2 | leaf_palm1_2 |
| PRJNA540215 | SRR9026567 | leaf primodia | palm1 mutant leaf primodia sample 1 | leaf_palm1_1 |
| PRJNA540215 | SRR9026568 | leaf primodia | wild type leaf primodia sample 2 | leaf_wd_2 |
| PRJNA540215 | SRR9026569 | leaf primodia | wild type leaf primodia sample 1 | leaf_wd_1 |
| PRJNA540215 | SRR9026570 | root | root | root |
| PRJNA657410 | SRR12458432 | root | Salt_27h | VR27_1.1 |
| PRJNA657410 | SRR12458433 | leaf | Salt_27h | HL27_1.1 |
| PRJNA657410 | SRR12458434 | root | Control | VR0_4.1 |
| PRJNA657410 | SRR12458435 | root | Control | VR0_3.1 |
| PRJNA657410 | SRR12458436 | root | Control | VR0_2.1 |
| PRJNA657410 | SRR12458437 | root | Control | VR0_1.1 |
| PRJNA657410 | SRR12458438 | leaf | Salt_3h | VL3_4.1 |
| PRJNA657410 | SRR12458439 | leaf | Salt_3h | VL3_3.1 |
| PRJNA657410 | SRR12458440 | leaf | Salt_3h | VL3_2.1 |
| PRJNA657410 | SRR12458441 | leaf | Salt_3h | VL3_1.1 |
| PRJNA657410 | SRR12458442 | leaf | Salt_27h | VL27_4.1 |
| PRJNA657410 | SRR12458443 | leaf | Salt_27h | VL27_3.1 |
| PRJNA657410 | SRR12458444 | leaf | Control | HL0_4.1 |
| PRJNA657410 | SRR12458445 | leaf | Salt_27h | VL27_2.1 |
| PRJNA657410 | SRR12458446 | leaf | Salt_27h | VL27_1.1 |
| PRJNA657410 | SRR12458447 | leaf | Control | VL0_4.1 |
| PRJNA657410 | SRR12458448 | leaf | Control | VL0_3.1 |
| PRJNA657410 | SRR12458449 | leaf | Control | VL0_2.1 |
| PRJNA657410 | SRR12458450 | leaf | Control | VL0_1.1 |
| PRJNA657410 | SRR12458451 | root | Salt_3h | HR3_4.1 |
| PRJNA657410 | SRR12458452 | root | Salt_3h | HR3_3.1 |
| PRJNA657410 | SRR12458453 | root | Salt_3h | HR3_2.1 |
| PRJNA657410 | SRR12458454 | root | Salt_3h | HR3_1.1 |
| PRJNA657410 | SRR12458455 | leaf | Control | HL0_3.1 |
| PRJNA657410 | SRR12458456 | root | Salt_27h | HR27_4.1 |
| PRJNA657410 | SRR12458457 | root | Salt_27h | HR27_3.1 |
| PRJNA657410 | SRR12458458 | root | Salt_27h | HR27_2.1 |
| PRJNA657410 | SRR12458459 | root | Salt_28h | HR27_1.1 |
| PRJNA657410 | SRR12458460 | root | Control | HR0_4.1 |
| PRJNA657410 | SRR12458461 | root | Control | HR0_3.1 |
| PRJNA657410 | SRR12458462 | root | Control | HR0_2.1 |
| PRJNA657410 | SRR12458463 | root | Control | HR0_1.1 |
| PRJNA657410 | SRR12458464 | root | Salt_3h | VR3_4.2 |
| PRJNA657410 | SRR12458465 | root | Salt_3h | VR3_3.2 |
| PRJNA657410 | SRR12458466 | root | Salt_3h | VR3_2.2 |
| PRJNA657410 | SRR12458467 | root | Salt_3h | VR3_1.2 |
| PRJNA657410 | SRR12458468 | root | Salt_27h | VR27_4.2 |
| PRJNA657410 | SRR12458469 | root | Salt_27h | VR27_3.2 |
| PRJNA657410 | SRR12458470 | leaf | Salt_3h | HL3_2.1 |
| PRJNA657410 | SRR12458471 | root | Salt_27h | VR27_2.2 |
| PRJNA657410 | SRR12458472 | root | Salt_27h | VR27_1.2 |
| PRJNA657410 | SRR12458473 | root | Control | VR0_4.2 |
| PRJNA657410 | SRR12458474 | root | Control | VR0_3.2 |
| PRJNA657410 | SRR12458475 | root | Control | VR0_2.2 |
| PRJNA657410 | SRR12458476 | root | Control | VR0_1.2 |
| PRJNA657410 | SRR12458477 | leaf | Salt_3h | VL3_4.2 |
| PRJNA657410 | SRR12458478 | leaf | Salt_3h | VL3_3.2 |
| PRJNA657410 | SRR12458479 | leaf | Salt_3h | VL3_2.2 |
| PRJNA657410 | SRR12458480 | leaf | Salt_3h | VL3_1.2 |
| PRJNA657410 | SRR12458481 | leaf | Salt_3h | HL3_1.1 |
| PRJNA657410 | SRR12458482 | leaf | Salt_27h | VL27_4.2 |
| PRJNA657410 | SRR12458483 | leaf | Salt_27h | VL27_3.2 |
| PRJNA657410 | SRR12458484 | leaf | Salt_27h | VL27_2.2 |
| PRJNA657410 | SRR12458485 | leaf | Salt_27h | VL27_1.2 |
| PRJNA657410 | SRR12458486 | leaf | Control | VL0_4.2 |
| PRJNA657410 | SRR12458487 | leaf | Control | VL0_3.2 |
| PRJNA657410 | SRR12458488 | leaf | Control | VL0_2.2 |
| PRJNA657410 | SRR12458489 | leaf | Control | VL0_1.2 |
| PRJNA657410 | SRR12458490 | root | Salt_3h | HR3_4.2 |
| PRJNA657410 | SRR12458491 | root | Salt_3h | HR3_3.2 |
| PRJNA657410 | SRR12458492 | leaf | Salt_3h | HL3_4.1 |
| PRJNA657410 | SRR12458493 | leaf | Salt_3h | HL3_3.1 |
| PRJNA657410 | SRR12458494 | leaf | Control | HL0_2.1 |
| PRJNA657410 | SRR12458495 | leaf | Control | HL0_1.1 |
| PRJNA657410 | SRR12458496 | leaf | Salt_27h | HL27_4.1 |
| PRJNA657410 | SRR12458497 | root | Salt_3h | HR3_2.2 |
| PRJNA657410 | SRR12458498 | root | Salt_3h | HR3_1.2 |
| PRJNA657410 | SRR12458499 | root | Salt_27h | HR27_4.2 |
| PRJNA657410 | SRR12458500 | root | Salt_27h | HR27_3.2 |
| PRJNA657410 | SRR12458501 | root | Salt_27h | HR27_2.2 |
| PRJNA657410 | SRR12458502 | root | Salt_27h | HR27_1.2 |
| PRJNA657410 | SRR12458503 | root | Control | HR0_4.2 |
| PRJNA657410 | SRR12458504 | root | Control | HR0_3.2 |
| PRJNA657410 | SRR12458505 | root | Control | HR0_2.2 |
| PRJNA657410 | SRR12458506 | root | Control | HR0_1.2 |
| PRJNA657410 | SRR12458507 | leaf | Salt_27h | HL27_3.1 |
| PRJNA657410 | SRR12458508 | leaf | Salt_3h | HL3_4.2 |
| PRJNA657410 | SRR12458509 | leaf | Salt_3h | HL3_3.2 |
| PRJNA657410 | SRR12458510 | leaf | Salt_3h | HL3_2.2 |
| PRJNA657410 | SRR12458511 | leaf | Salt_3h | HL3_1.2 |
| PRJNA657410 | SRR12458512 | leaf | Salt_27h | HL27_4.2 |
| PRJNA657410 | SRR12458513 | leaf | Salt_27h | HL27_3.2 |
| PRJNA657410 | SRR12458514 | leaf | Salt_27h | HL27_2.2 |
| PRJNA657410 | SRR12458515 | leaf | Salt_27h | HL27_1.2 |
| PRJNA657410 | SRR12458516 | leaf | Control | HL0_4.2 |
| PRJNA657410 | SRR12458517 | leaf | Control | HL0_3.2 |
| PRJNA657410 | SRR12458518 | leaf | Salt_27h | HL27_2.1 |
| PRJNA657410 | SRR12458519 | leaf | Control | HL0_2.2 |
| PRJNA657410 | SRR12458520 | leaf | Control | HL0_1.2 |
| PRJNA657410 | SRR12458521 | root | Salt_3h | VR3_4.1 |
| PRJNA657410 | SRR12458522 | root | Salt_3h | VR3_3.1 |
| PRJNA657410 | SRR12458523 | root | Salt_3h | VR3_2.1 |
| PRJNA657410 | SRR12458524 | root | Salt_3h | VR3_1.1 |
| PRJNA657410 | SRR12458525 | root | Salt_27h | VR27_4.1 |
| PRJNA657410 | SRR12458526 | root | Salt_27h | VR27_3.1 |
| PRJNA657410 | SRR12458527 | root | Salt_27h | VR27_2.1 |
| PRJNA742658 | SRR14999926 | seedling | 137m NaCl+MT | MT2 |
| PRJNA742658 | SRR14999927 | seedling | 137m NaCl+MT | MT1 |
| PRJNA742658 | SRR14999928 | seedling | control | CK3 |
| PRJNA742658 | SRR14999929 | seedling | control | CK2 |
| PRJNA742658 | SRR14999930 | seedling | control | CK1 |
| PRJNA742658 | SRR14999931 | seedling | 137m NaCl | Salt3 |
| PRJNA742658 | SRR14999932 | seedling | 137m NaCl | Salt2 |
| PRJNA742658 | SRR14999933 | seedling | 137m NaCl | Salt1 |
| PRJNA742658 | SRR14999934 | seedling | 137m NaCl+MT | MT3 |
Laboratory: Key Laboratory of Tropical Plant Resources and Sustainable Use
Institute: Xishuangbanna Tropical Botanical Garden, Chinese Academy of Sciences.
Address: No. 21 Qingsong Road, Ciba, Kunming 650204, Yunnan Province, China
E-mail: liuchangning@xtbg.ac.cn